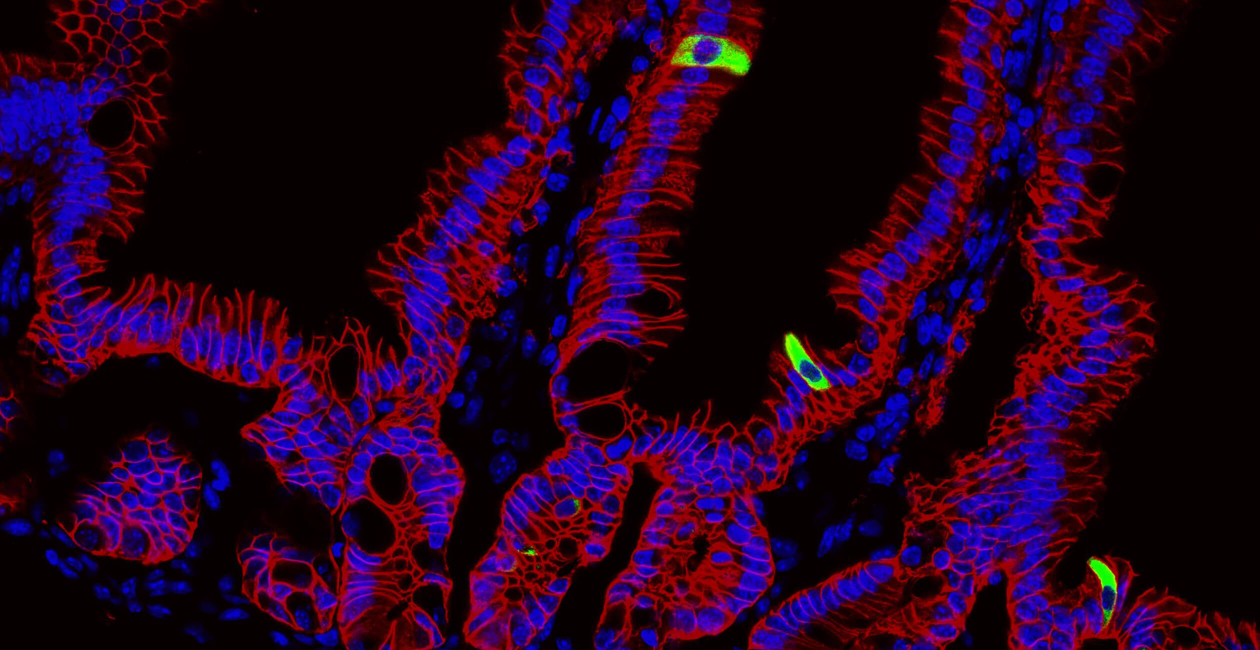
The Harrison Lab studies the mechanisms controlling host-microbe interactions at barrier tissues, primarily the skin and the gut. We perform our research in a multidisciplinary and collaborative manner, combining in vivo cellular and molecular immunology, using genetic mouse models, microbiology, transcriptomic, and epigenetic analyses, to understand how commensal-specific immunity contributes to tissue homeostasis and repair.
A major research interest in the lab is studying the role of commensal-specific T and B cells in the skin and gastrointestinal tract. To do so, we have generated new reagents, commensal-specific T and B cell tetramers, and T cell receptor transgenic mice, to enable us to identify, profile, and manipulate commensal-specific immune responses following commensal colonization, and during experimental infection and injury.
Our goal is to understand how these immune cells promote barrier tissue integrity and repair, and to understand how this goes awry during disease.

Oliver Harrison, DPhil
Lab Members

Addison Gralen

Hannah Kalinoski, PhD

Tayla Olsen

Sarah Pemberton

Sheenam Verma, PhD
Research Projects
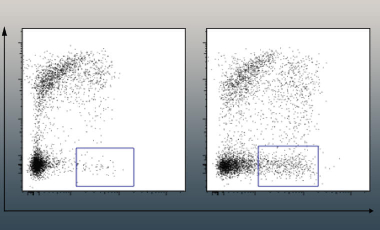
Commensal-specific B cells
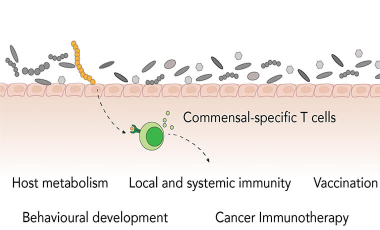
Commensal-Specific T cells
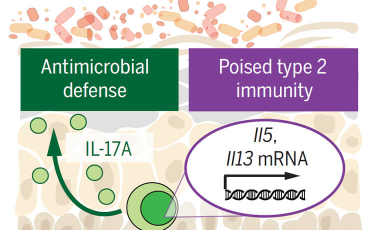
Poised homeostatic immunity
Featured Publications
-
Jul 2020
Poised for tissue repair.
ScienceHarrison OJ -
Jan 2019
Commensal-specific T cell plasticity promotes rapid tissue adaptation to injury.
ScienceHarrison OJ, Linehan JL, Shih HY, Bouladoux N, Han SJ, Smelkinson M, Sen SK, Byrd AL, Enamorado M, Yao C, Tamoutounour S, Van Laethem F, Hurabielle C, Collins N, Paun A, Salcedo R, O'Shea JJ, Belkaid Y -
Feb 2018
c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont.
NatureXu M, Pokrovskii M, Ding Y, Yi R, Au C, Harrison OJ, Galan C, Belkaid Y, Bonneau R, Littman DR -
Jan 2018
Non-classical Immunity Controls Microbiota Impact on Skin Immunity and Tissue Repair.
CellLinehan JL, Harrison OJ, Han SJ, Byrd AL, Vujkovic-Cvijin I, Villarino AV, Sen SK, Shaik J, Smelkinson M, Tamoutounour S, Collins N, Bouladoux N, Dzutsev A, Rosshart SP, Arbuckle JH, Wang CR, Kristie TM, Rehermann B, Trinchieri G, Brenchley JM, O'Shea JJ, Belkaid Y -
Apr 2017
Homeostatic Immunity and the Microbiota.
ImmunityBelkaid Y, Harrison OJ