First Study to Investigate the Human Genome in Multiple Sclerosis
Benaroya Research Institute Receives $1.9 Million from National Institutes of Health
The National Institutes of Health (NIH) awarded Benaroya Research Institute at Virginia Mason (BRI) a $1.9 million grant to find marks in the human genome which can explain why some white blood cells cause damage to the spinal cord and brain in multiple sclerosis (MS). This is the first study to look for molecular changes in the genome of specific immune cells responsible for the devastation caused by MS. The broad-based study will determine the function of these cells, how they are generated and how they can be regulated in system models of MS and in humans.
"We want to understand the factors that make these cells target the spinal cord and brain to cause disease," says Estelle Bettelli, PhD, BRI Assistant Member and co-principal investigator of the study. Dr. Bettelli and other scientists have identified different types of T cells which they believe are potent inducers of MS and other autoimmune diseases. She has also developed system models to study different forms of multiple sclerosis.
"With Dr. Bettelli's research advances and with the new technological innovations in genome research, we can look at specific marks present in the genome of these cells and understand how they are generated and how they can be controlled," says co-principal investigator Steven Ziegler, PhD, Director of the BRI Immunology Research Program. Dr. Ziegler has used whole genome studies to investigate these cell types in healthy individuals. "We can then see how the genomic marks affect the cells in model systems of MS and how they operate in humans cells with and without the disease. We can also see how these cells behave once the patient receives treatment and if various treatments make the cells act differently."
"It is important to know how and when these cells are formed in the body to determine how to inhibit their harmful function," says Dr. Bettelli. "It is becoming clear that MS is not a unique disease entity but can present itself in different clinical forms and variants. Several factors, including the cell types involved, are believed to dictate the clinical progression of MS. The understanding of how and which cell populations of the immune system participate in the autoimmune attack is very important for determining current treatments and designing new therapeutics tailored to the different forms of MS. We hope to find ways to significantly inhibit these dangerous cells with new targeted medicines with fewer side effects.
"This work highlights a key mechanism for understanding and modifying the immune cells that cause autoimmune diseases like MS," says BRI Director Gerald Nepom, MD, PhD. "It is an exciting example of the power of merging new genomic technologies with state-of-the-art immunology research to address a major clinical need."
Other scientists collaborating in this effort are Jane Buckner, MD, BRI Associate Director, Damien Chaussabel, PhD, BRI Director of Systems Immunology, Mariko Kita, MD, BRI Affiliate Investigator and Director of the Virginia Mason Multiple Sclerosis Center, and John Stamatoyannopoulos, MD, Associate Professor of Genome Sciences and Medicine, University of Washington.
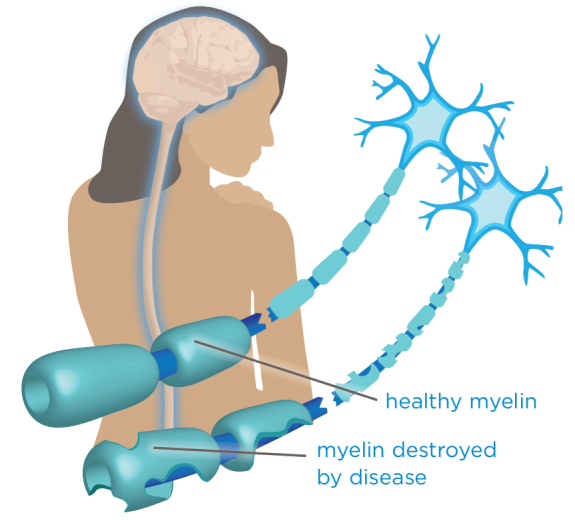
Multiple sclerosis is an autoimmune disease in which the body's immune system mistakenly attacks myelin, the fatty substance that surrounds and protects the nerve fibers in the central nervous system. When the myelin is damaged the nerve impulses are not transmitted as quickly or efficiently, resulting in symptoms such as numbness in the limbs, fatigue, dizziness, paralysis and/or loss of vision. Symptoms of MS will often improve and relapse with time and vary from one person to another. In progressive forms of multiple sclerosis, they gradually worsen.
MS affects approximately 400,000 Americans but is much more common in the Northwest. Some likely factors that contribute to this may be vitamin D deficiency, genetic predisposition and environmental triggers. Other factors are still unknown.


